Difference between revisions of "MAPK6-DT"
(Created page with "''MAPK6-DT'', a divergent lncRNA (long noncoding RNA) of MAPK6 overexpressed in liver tumorigenesis. ==Annotated Information== ===Name=== ''MAPK6-DT'': MAPK6 divergent trans...") |
(→Expression) |
||
| Line 17: | Line 17: | ||
===Expression=== | ===Expression=== | ||
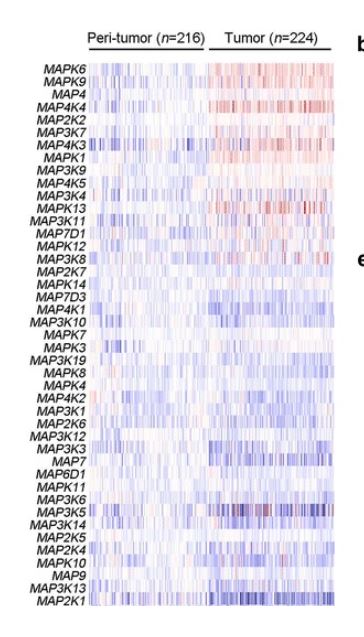
| − | lncMAPK6 overexpression through CRISPRa strategy promoted the self-renewal of liver TICs.<ref name="ref1" /> | + | [[File: MAPK6-DT.JPG|right|thumb|400px|'''Overexpression of MAPK6 in liver tumor and TICs. Expression profiles of MAPK components were shown as heatmap. Blue denoted low expression and red denoted high expression. ''' <ref name="ref1" />.]] |
| + | lncMAPK6 overexpression through CRISPRa strategy promoted the self-renewal of liver TICs.<ref name="ref1" /> | ||
| + | |||
===Diseases=== | ===Diseases=== | ||
Liver cancer <ref name="ref1" /> | Liver cancer <ref name="ref1" /> | ||
Revision as of 07:20, 18 December 2018
MAPK6-DT, a divergent lncRNA (long noncoding RNA) of MAPK6 overexpressed in liver tumorigenesis.
Contents
Annotated Information
Name
MAPK6-DT: MAPK6 divergent transcript (HGNC:53900)
Alias
lncMAPK6
Characteristics
MAPK6-DT is a divergent lncRNA (long noncoding RNA) of MAPK6 with chromosomal location at 15q21.2. Divergent lncRNAs, often modulate their nearby genes in cis, are positively-related to their nearby genes. As a divergent lncRNA of MAPK6, lncMAPK6 is co-expressed with MAPK6 in liver tumor and TICs.
Function
LncMAPK6 regulates the expression of MAPK6 through transcriptional activation. LncMAPK6 promotes liver tumor propagation and TIC self-renewal through MAPK6. LncMAPK6 interacts with and recruits RNA polymerase II to MAPK6 promoter, and finally activates the transcription of MAPK6. Through MAPK6 transcriptional regulation, lncMAPK6 drives MARK signaling activation.
Regulation
MAPK6 is critical for the function of lncMAPK6 as after MAPK6 knockout, lncMAPK6 had no effect on liver TIC self-renewal. LncMAPK6 had no effect on tumor invasion if MAPK6 was knockout, further confirming the importance of MAPK6 in lncMAPK6 signaling
Expression
lncMAPK6 overexpression through CRISPRa strategy promoted the self-renewal of liver TICs.[1]
Diseases
Liver cancer [1]
Labs working on this lncRNA
- Department of general surgery, The Fifth Affiliated Hospital of Guangzhou Medical University, Guangdong Sheng, China
- Department of Abdominal Oncology, The Fifth Affiliated Hospital of Guangzhou Medical University, Guangdong Sheng, China
- Department of General Surgery, Guangdong General Hospital, Guangdong Academy of Medical Sciences, Guangdong Sheng, China
References
Sequence
>gi|112543478|ref|NR_156732.1| Homo sapiens MAPK6 divergent transcript (MAPK6-DT), long non-coding RNA
000081 GGAAGACAGC TGGGGCCGTT TGCTCTACTA CCTGCCCTGG CATTAAGGGA ACAAAGTGAC CAAACCCAGT GCCAGCGTTT 000160
000161 ACTAAGCACC TACTACGTGC AAGGCACTGG GCAATCCGAG AGGAAGAGGC CTCAGTCTCC AGGAAGGGTG GACCTTAAGA 000240
000241 TGGATTGGCT GGAAGGCGAC AGCCCGAGCA CAAGACAGAC GCGTTAGAGG CCAATACTAA GGTCATTCAT TCTGCCAGAG 000320
000321 TATGGAGGAA GGCTTTCAGG AGTGGTCGCC AAGGTGGCCT TTCAGCTGGA TCCCGGAGGA GCGGATATAA CCAGGAGGGC 000400
000401 ACTTCACAGC CTTCTGTGCT TAGGAACTCT ATACAGCACT TCAGCAATAG ATTTGGGGGC CACTTTAAAC AGCAAAATCA 000480
000481 CCAACGGAAA GCAAAACATG TGATGGCTCG GCACAGTGGC ATGTGCCTAT AGTCCCAGCT ACTCAGGAGG CTGAGACAGG 000560
000561 AGGGTCACTT GAGCCCAGGA GTTTGAGTCC AGCCTGGTCA ACATAGAGAG ATCCTGTTTC TAAAAACAAA AAAATCATGG 000640
000641 CACCAAATAG ACCATGGAAA GGACACTCGT TTACAGTACG AGAGTTGAAA CAAGAAGGCA GAATCTTGTC TTGTTTGGCC 000720
000721 TCAGCTGGCA ACGTTCCCGT AGGGTGACTC AAGTATCTCG TTACTGTGCA TGTTCAAAAG TGACTACAAA GGTGCTATGA 000800
000801 CTACTTCGAG GTTACAAATA AGTTTCAGCA ATAGGCAAAT TTGCAAACAG AATCTGTGAA TAATG